Problem setting: persistence for big data
Joblib is a powerful Python package for management of computation: parallel computing, caching, and primitives for out-of-core computing. It is handy when working on so called big data, that can consume more than the available RAM (several GB nowadays). In such situations, objects in the working space must be persisted to disk, for out-of-core computing, distribution of jobs, or caching.
An efficient strategy to write code dealing with big data is to rely on numpy arrays to hold large chunks of structured data. The code then handles objects or arbitrary containers (list, dict) with numpy arrays. For data management, joblib provides transparent disk persistence that is very efficient with such objects. The internal mechanism relies on specializing pickle to handle better numpy arrays.
Recent improvements reduce vastly the memory overhead of data persistence.
Limitations of the old implementation
❶ Dumping/loading persisted data with compression was a memory hog, because of internal copies of data, limiting the maximum size of usable data with compressed persistence:
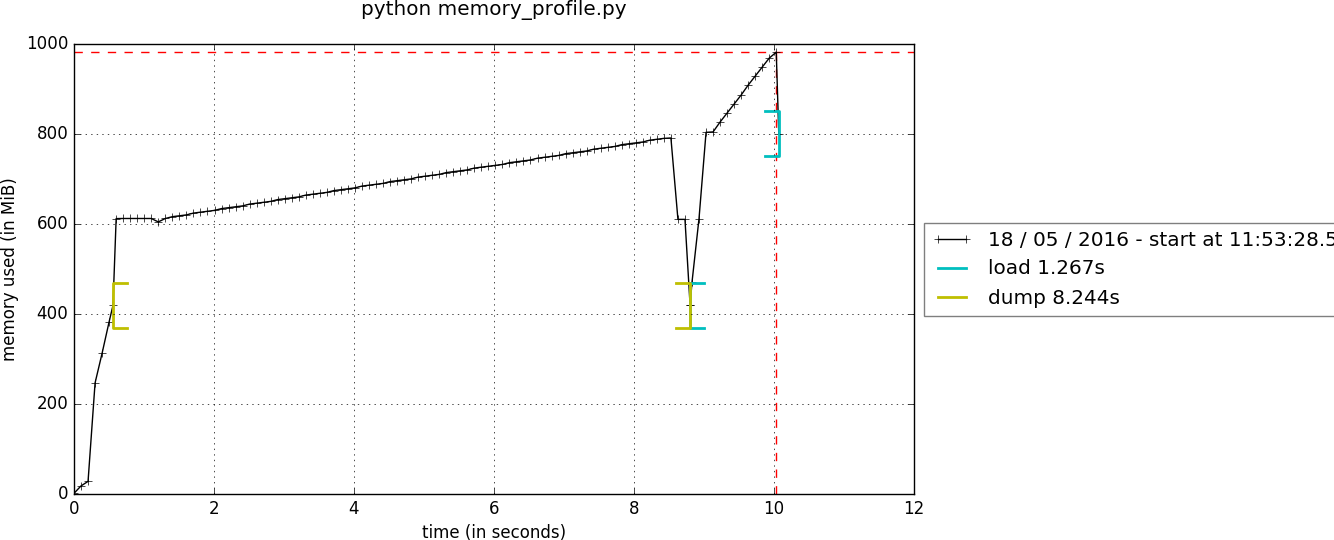
We see the increased memory usage during the calls to dump and load functions, profiled using the memory_profiler package with this gist
❷ Another drawback was that large numpy arrays (>10MB) contained in an arbitrary Python object were dumped in separate .npy file, increasing the load on the file system [1]:
>>> import numpy as np
>>> import joblib # joblib version: 0.9.4
>>> obj = [np.ones((5000, 5000)), np.random.random((5000, 5000))]
# 3 files are generated:
>>> joblib.dump(obj, '/tmp/test.pkl', compress=True)
['/tmp/test.pkl', '/tmp/test.pkl_01.npy.z', '/tmp/test.pkl_02.npy.z']
>>> joblib.load('/tmp/test.pkl')
[array([[ 1., 1., ..., 1., 1.]],
array([[ 0.47006195, 0.5436392 , ..., 0.1218267 , 0.48592789]])]
What’s new: compression, low memory…
❶ Memory usage is now stable:
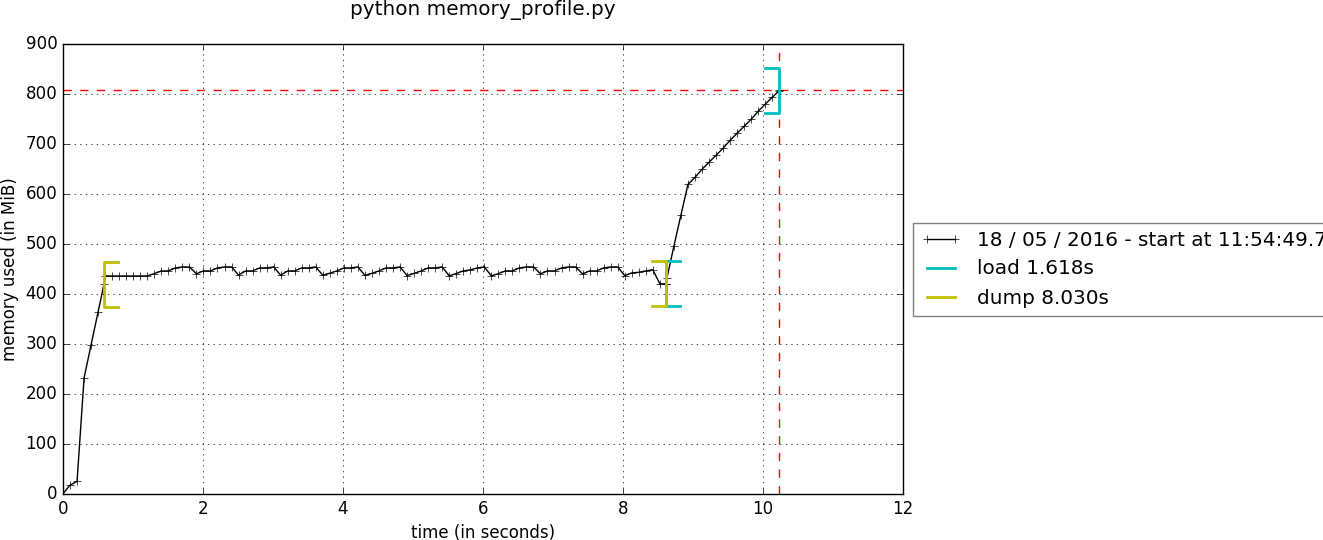
❷ All numpy arrays are persisted in a single file:
>>> import numpy as np
>>> import joblib # joblib version: 0.10.0 (dev)
>>> obj = [np.ones((5000, 5000)), np.random.random((5000, 5000))]
# only 1 file is generated:
>>> joblib.dump(obj, '/tmp/test.pkl', compress=True)
['/tmp/test.pkl']
>>> joblib.load('/tmp/test.pkl')
[array([[ 1., 1., ..., 1., 1.]],
array([[ 0.47006195, 0.5436392 , ..., 0.1218267 , 0.48592789]])]
❸ Persistence in a file handle (ongoing work in a pull request)
❹ More compression formats are available
Backward compatibility
Existing joblib users can be reassured: the new version is still compatible with pickles generated by older versions (>= 0.8.4). You are encouraged to update (rebuild?) your cache if you want to take advantage of this new version.
Benchmarks: speed and memory consumption
Joblib strives to have minimum dependencies (only numpy) and to be agnostic to the input data. Hence the goals are to deal with any kind of data while trying to be as efficient as possible with numpy arrays.
To illustrate the benefits and cost of the new persistence implementation, let’s now compare a real life use case (LFW dataset from scikit-learn) with different libraries:
- Joblib, with 2 different versions, 0.9.4 and master (dev),
- Pickle
- Numpy
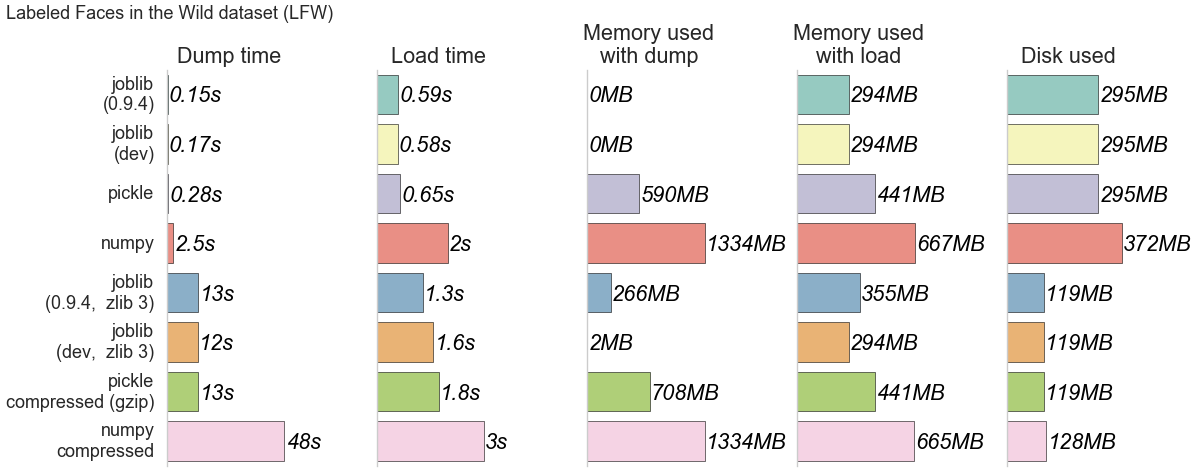
The four first lines use non compressed persistence strategies, the last four use persistence with zlib/gzip [2] strategies. Code to reproduce the benchmarks is available on this gist.
⚫ Speed: the results between joblib 0.9.4 and 0.10.0 (dev) are similar whereas numpy and pickle are clearly slower than joblib in both compressed and non compressed cases.
⚫ Memory consumption: Without compression, old and new joblib versions are the same; with compression, the new joblib version is much better than the old one. Joblib clearly outperforms pickle and numpy in terms of memory consumption. This can be explained by the fact that numpy relies on pickle if the object is not a pure numpy array (a list or a dict with arrays for example), so in this case it inherits the memory drawbacks from pickle. When persisting pure numpy arrays (not tested here), numpy uses its internal save/load functions which are efficient in terms of speed and memory consumption.
⚫ Disk used: results are as expected: non compressed files have the same size as the in-memory data; compressed files are smaller.
Caveat Emptor: performance is data-dependent
Different data compress more or less easily. Speed and disk used will vary depending on the data. Key considerations are:
- Fraction of data in arrays: joblib is efficient if much of the data is contained in numpy arrays. The worst case scenario is something like a large dictionary of random numbers as keys and values.
- Entropy of the data: an array fully of zeros will compress well and fast. A fully random array will compress slowly, and use a lot of disk. Real data is often somewhere in the middle.
Extra improvements in compressed persistence
New compression formats
Joblib can use new compression formats based on Python standard library modules: zlib, gzip, bz2, lzma and xz (the last 2 are available for Python greater than 3.3). The compressor is selected automatically when the file name has an explicit extension:
>>> joblib.dump(obj, '/tmp/test.pkl.z') # zlib
['/tmp/test.pkl.z']
>>> joblib.dump(obj, '/tmp/test.pkl.gz') # gzip
['/tmp/test.pkl.gz']
>>> joblib.dump(obj, '/tmp/test.pkl.bz2') # bz2
['/tmp/test.pkl.bz2']
>>> joblib.dump(obj, '/tmp/test.pkl.lzma') # lzma
['/tmp/test.pkl.lzma']
>>> joblib.dump(obj, '/tmp/test.pkl.xz') # xz
['/tmp/test.pkl.xz']
One can tune the compression level, setting the compressor explicitly:
>>> joblib.dump(obj, '/tmp/test.pkl.compressed', compress=('zlib', 6))
['/tmp/test.pkl.compressed']
>>> joblib.dump(obj, '/tmp/test.compressed', compress=('lzma', 6))
['/tmp/test.pkl.compressed']
On loading, joblib uses the magic number of the file to determine the right decompression method. This makes loading compressed pickle transparent:
>>> joblib.load('/tmp/test.compressed')
[array([[ 1., 1., ..., 1., 1.]],
array([[ 0.47006195, 0.5436392 , ..., 0.1218267 , 0.48592789]])]
Importantly, the generated compressed files use a standard compression file format: for instance, regular command line tools (zip/unzip, gzip/gunzip, bzip2, lzma, xz) can be used to compress/uncompress a pickled file generated with joblib. Joblib will be able to load cache compressed with those tools.
Toward more and faster compression
Specific compression strategies have been developped for fast compression, sometimes even faster than disk reads such as snappy , blosc, LZO or LZ4. With a file-like interface, they should be readily usable with joblib.
In the benchmarks above, loading and dumping with compression is slower than without (though only by a factor of 3 for loading). These were done on a computer with an SSD, hence with very fast I/O. In a situation with slower I/O, as on a network drive, compression could save time. With faster compressors, compression will save time on most hardware.
Compressed persistence into a file handle
Now that everything is stored in a single file using standard compression formats, joblib can persist in an open file handle:
>>> with open('/tmp/test.pkl', 'wb') as f:
>>> joblib.dump(obj, f)
['/tmp/test.pkl']
>>> with open('/tmp/test.pkl', 'rb') as f:
>>> print(joblib.load(f))
[array([[ 1., 1., ..., 1., 1.]],
array([[ 0.47006195, 0.5436392 , ..., 0.1218267 , 0.48592789]])]
This also works with compression file object available in the standard library, like gzip.GzipFile, bz2.Bz2File or lzma.LzmaFile:
>>> import gzip
>>> with gzip.GzipFile('/tmp/test.pkl.gz', 'wb') as f:
>>> joblib.dump(data, f)
['/tmp/test.pkl.gz']
>>> with gzip.GzipFile('/tmp/test.pkl.gz', 'rb') as f:
>>> print(joblib.load(f))
[array([[ 1., 1., ..., 1., 1.]],
array([[ 0.47006195, 0.5436392 , ..., 0.1218267 , 0.48592789]])]
Be sure that you use a decompressor matching the internal compression when loading with the above method. If unsure, simply use open, joblib will select the right decompressor:
>>> with open('/tmp/test.pkl.gz', 'rb') as f:
>>> print(joblib.load(f))
[array([[ 1., 1., ..., 1., 1.]],
array([[ 0.47006195, 0.5436392 , ..., 0.1218267 , 0.48592789]])]
Towards dumping to elaborate stores
Working with file handles opens the door to storing cache data in database blob or cloud storage such as Amazon S3, Amazon Glacier and Google Cloud Storage (for instance via the Python package boto).
Implementation
A Pickle Subclass: joblib relies on subclassing the Python Pickler/Unpickler [3]. These are state machines that walk the graph of nested objects (a dict may contain a list, that may contain…), creating a string representation of each object encountered. The new implementation proceeds as follows:
- Pickling an arbitrary object: when an np.ndarray object is reached, instead of using the default pickling functions (__reduce__()), the joblib Pickler replaces in pickle stream the ndarray with a wrapper object containing all important array metadata (shape, dtype, flags). Then it writes the array content in the pickle file. Note that this step breaks the pickle compatibility. One benefit is that it enables using fast code for copyless handling of the numpy array. For compression, we pass chunks of the data to a compressor object (using the buffer protocol to avoid copies).
- Unpickling from a file: when pickle reaches the array wrapper, as the object is in the pickle stream, the file handle is at the beginning of the array content. So at this point the Unpickler simply constructs an array based on the metadata contained in the wrapper and then fills the array buffer directly from the file. The object returned is the reconstructed array, the array wrapper being dropped. A benefit is that if the data is stored not compressed, the array can be directly memory mapped from the storage (the mmap_mode option of joblib.load).
This technique allows joblib to pickle all objects in a single file but also to have memory-efficient dump and load.
A fast compression stream: as the pickling refactoring opens the door to file objects usage, joblib is now able to persist data in any kind of file object: open, gzip.GzipFile, bz2.Bz2file and lzma.LzmaFile. For performance reason and usability, the new joblib version uses its own file object BinaryZlibFile for zlib compression. Compared to GzipFile, it disables crc computation, which bring a performance gain of 15%.
Speed penalties of on-the-fly writes
There’s also a small speed difference with dict/list objects between new/old joblib when using compression. The old version pickles the data inside a io.BytesIO buffer and then compress it in a row whereas the new version write “on the fly” compressed chunk of pickled data to the file. Because of this internal buffer the old implementation is not memory safe as it indeed copy the data in memory before compressing. The small speed difference was judged acceptable compared to this memory duplication.
Conclusion and future work
Memory copies were a limitation when caching on disk very large numpy arrays, e.g arrays with a size close to the available RAM on the computer. The problem was solved via intensive buffering and a lot of hacking on top of pickle and numpy. Unfortunately, our strategy has poor performance with big dictionaries or list compared to a cPickle, hence try to use numpy arrays in your internal data structures (note that something like scipy sparse matrices works well, as it builds on arrays).
For the future, maybe numpy’s pickle methods could be improved and make a better use of 64-bit opcodes for large objects that were introduced in Python recently.
Pickling using file handles is a first step toward pickling in sockets, enabling broadcasting of data between computing units on a network. This will be priceless with joblib’s new distributed backends.
Other improvements will come from better compressor, making everything faster.
Note
The pull request was implemented by @aabadie. He thanks @lesteve, @ogrisel and @GaelVaroquaux for the valuable help, reviews and support.
| [1] | The load created by multiple files on the filesystem is particularly detrimental for network filesystems, as it triggers multiple requests and isn’t cache friendly. |
| [2] | gzip is based on zlib with additional crc checks and a default compression level of 3. |
| [3] | A drawback of subclassing the Python Pickler/Unpickler is that it is done for the pure-Python version, and not the “cPickle” version. The latter is much faster when dealing with a large number of Python objects. Once again, joblib is efficient when most of the data is represented as numpy arrays or subclasses. |
